 Aaron Pleitner
Aaron Pleitner
 Aaron Pleitner
Aaron Pleitner
Are you struggling to find the root cause of repeat positives in your food facility’s production environment? Wasting time and money directing sanitation efforts only to keep getting positives? Feeling pressure to get a production line back up and running safely?
Microbial strain typing can help. It’s a powerful tool that food safety professionals use to help identify the root cause of frustrating repeat positives, conduct efficient traceback with informed seek & destroy efforts, and return to producing food products safely.
In this post, you will discover how strain typing differentiates between microbial isolates of pathogens like Listeria, Salmonella, and E. coli, and how it can help you identify the root cause of repeat positives so you can target your remediation efforts accordingly and take your environmental monitoring program to the next level.
Let’s get started.
What is Microbial Strain Typing?
Microbial strain typing is a laboratory method used to identify and differentiate among microbial isolates.
Several technologies exist that can identify species of microorganisms. However, there are several cases (such as when you get repeat positives) where it is helpful to classify organisms beyond the species level and discover whether two isolates of the same species are the same or different strain.
This is where strain typing enters the picture.
Microorganisms within the same species (for example, Listeria monocytogenes and E. coli) can have minor genetic differences, leading to different “strains” with varied characteristics and sources (for example, one L. monocytogenes strain may persist in a slicer harborage site while another appears only transiently from incoming raw materials; one E. coli strain may be a routine indicator organism while another is a Shiga toxin–producing strain such as E. coli O157:H7).
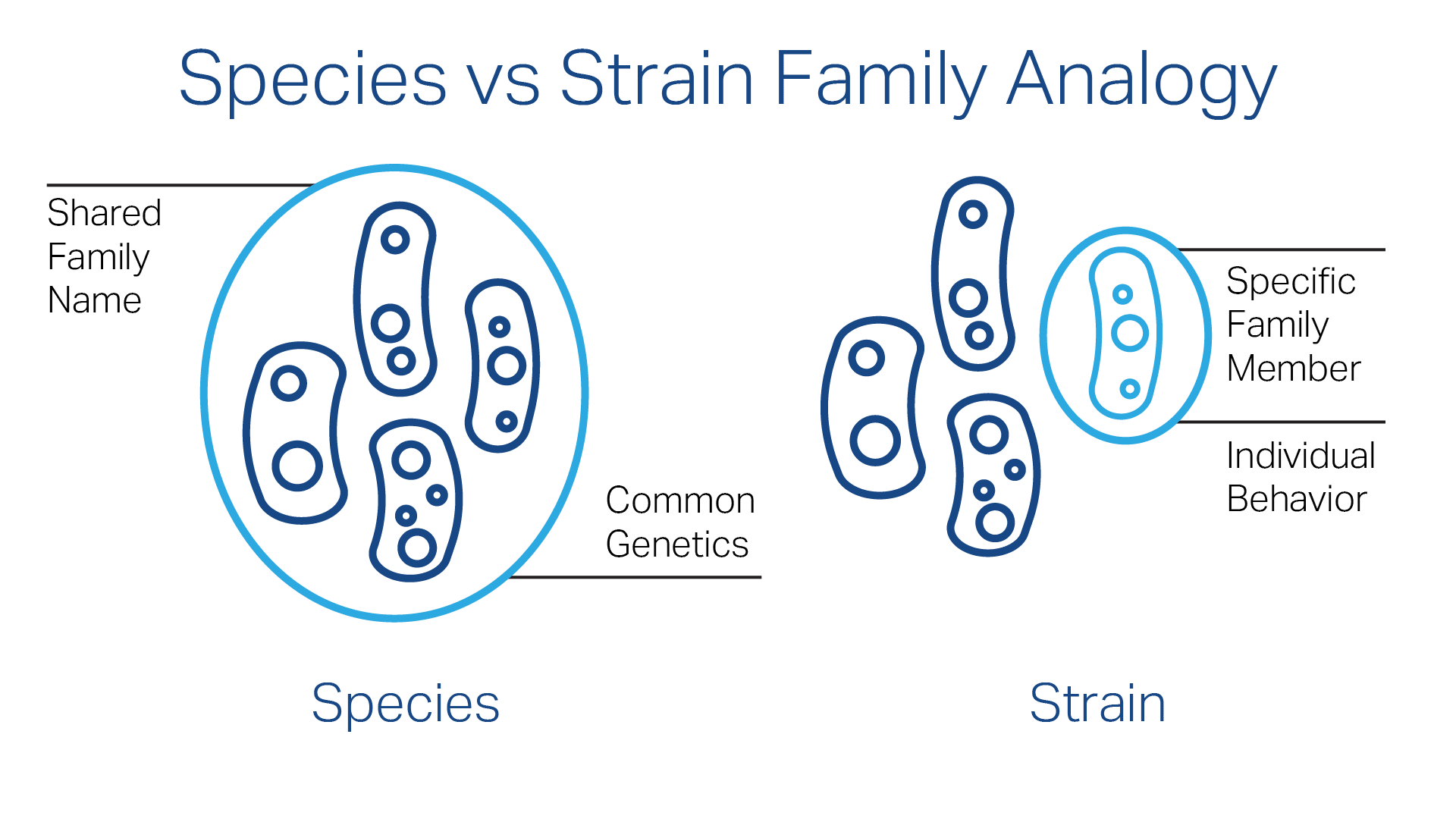
A useful analogy is to think of a species as a family’s shared last name, and individual strains as specific members of that family. They share common characteristics but remain distinguishable individuals.
Strain typing helps distinguish between these strains, aiding in the understanding of diverse microbial populations.
Why Perform Microbial Source Tracking?
So, how can strain typing help your organization produce food products more safely and in a cost-effective manner?
There are multiple reasons to track a particular microorganism through the food production process, but we’ll look at three of the most common and helpful reasons:
Identify a Pathogen Source
This is one of the most helpful uses of microbial strain typing. If you continue to get repeat positives from your production environment, you need to quickly identify the source so you can direct your remediation efforts effectively.
Otherwise, you could end up wasting time and money chasing the source without a complete picture of your target.
Strain typing helps decipher the origin of the pathogen and if it is persisting in your environment. It can tell you if the pathogen is transient and originates from your raw materials, personnel, or another source, or whether it’s coming from a harborage site in your facility.
This is critical information for uncovering the root cause and solving the problem for good.

Identify a Spoilage Source
Maybe your product is spoiling prior to reaching the end of its normal shelf-life. Microbial strain typing helps identify the source of spoilage organisms, such as unsanitary equipment or contaminated raw materials.
This information helps you target and eliminate the source of the spoilage organisms so your products can achieve their maximum shelf-life. This, of course, helps you control costs while protecting your brand reputation.
Track a Probiotic Through Your Process
There are also good microorganisms you may want to source-track, such as probiotics. Microbial strain typing allows you to determine if a probiotic starter culture is present in the finished product after packaging, providing insight into the effectiveness of your process.
The Science Behind Microbial Strain Typing
Now, let’s look at the primary methods laboratories use to perform strain typing. Each has benefits and potential drawbacks for food product manufacturers, so it’s important to choose the right method to suit your goals.
Whole Genome Sequencing (WGS)
This is the gold standard for microbial strain typing. It’s the same technology that was used to sequence the human genome, so it must be right for you, correct?
Not necessarily.
WGS identifies the complete DNA sequence of an organism’s genome at a single time. It also allows for the detection of all types of genetic variation in a single experiment, providing the highest resolution for strain differentiation.
For food product manufacturers, however, WGS has significant drawbacks:
- Very expensive.
- Takes several days to weeks to complete.
- Identifies the species of the isolate, which is an issue if you identify a pathogen.
The third bullet point is important for food product manufacturers, so let’s take a closer look.
Understanding the entire genetic makeup of an isolate and its species allows that information to be tied to regulatory agency databases (FDA, USDA, CDC), which can link you to a foodborne illness outbreak.
Additionally, identifying the species can be an issue if you identify a pathogen and your products are already in commerce – this situation could trigger a recall. Furthermore, this information can be tied to other isolates, revealing all the virulence factors and antibiotic-resistance profiles.
For these reasons, many food product manufacturers choose not to use WGS even though it’s considered the gold standard of strain typing.
Pulsed-Field Gel Electrophoresis (PFGE)
PFGE is a tried-and-true method of strain typing that has been around for many years.
It separates large DNA molecules by applying an electrical field through a sample of DNA suspended in place. The DNA, cut into fragments by specific restriction enzymes, migrates differently depending on its size and shape.
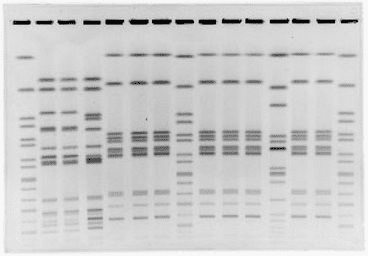
The resulting pattern – similar to a barcode (see image below) – is unique to each strain, enabling differentiation. It’s relatively inexpensive, but it takes several days to get results. Also, it can be used with a limited number of organisms.
Importantly, like Whole Genome Sequencing, it requires identifying the species of the microorganism, which has drawbacks as we have discussed.
Ribotyping
Ribotyping is a genetic “fingerprinting” method used to identify and differentiate bacterial species and strains. It targets the genes that code for the 16S ribosomal RNA in bacteria.
These genes are highly conserved, meaning they remain largely unchanged through evolution. However, they contain certain hypervariable regions that can differ significantly between different strains.
A ribotyping procedure involves extracting DNA, amplifying the 16S rRNA genes, and then comparing the patterns of these genes using a barcode, similar to PFGE.
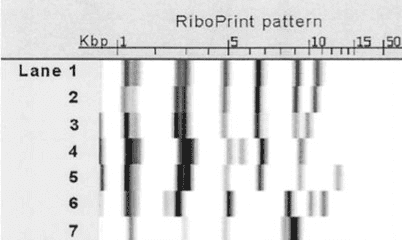
Ribotyping is a faster, relatively less expensive method compared to WGS and PFGE. However, it can also inadvertently identify the species of the microorganism during the process.
Phenotypic Fingerprinting
So, how can food product manufacturers perform microbial strain typing in a way that doesn’t identify the species but still provides detailed information at the subspecies level? That leads us to the next method – phenotypic fingerprinting, specifically using the Bruker IR Biotyper®.
This method involves the use of infrared spectroscopy to produce a unique spectral fingerprint of a microbial strain based on its phenotypic properties. Phenotype refers to the observable physical or biochemical characteristics of an organism.
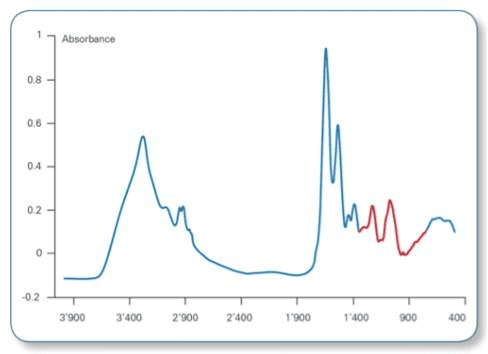
This allows for the comparison of multiple isolates without identifying the species, which offers benefits to food manufacturers, as we explained earlier.
For example, in a ready-to-eat facility, this technology allows you to make comparisons for “Listeria-like” organisms without identifying the species. This is critical for regulatory purposes since Listeria spp. may not implicate a product while Listeria monocytogenes will.
So, how does phenotypic fingerprinting with the Bruker IR Biotype work?
The data is analyzed with hierarchical cluster analysis (HCA) and results are displayed as a dendrogram or distance matrix. This allows you to see clusters of isolates and determine relatedness. This is the key to the analysis and allows you to make comparisons to determine if any isolates are related.
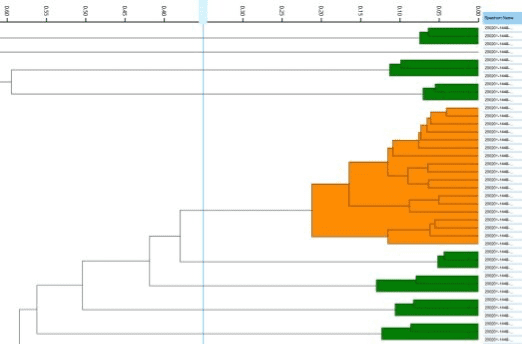
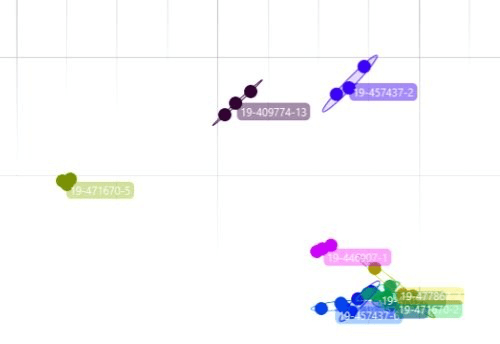
How Strain Typing Data Helps You
Now that you have all this data generated through strain typing, what do you do with it?
- Visualize the relatedness of positives, helping you direct your sanitation efforts effectively and eliminate the root cause of the problem. This can be added to any heat map profiling done with EMP data.
- Determine if isolates from a major contamination event are related (e.g. an event day for a slaughter facility).
- Determine if spoilage issues in final products are originating from a particular raw material or manufacturing harborage area without having to identify the organisms.
The Bruker IR Biotyper® method of microbial strain typing offers several advantages over the other methods discussed, which is why we frequently recommend it to our clients. In addition to providing information at the subspecies level without identifying the species, it produces results in a matter of hoursand is relatively less expensive than other methods. This enables you to run more isolates.
In fact, one beef slaughter plant ran 68 isolates in under 6 hours using this technology, which helped them identify the source of E. coli O157:H7 in their plant so they could target their decontamination efforts and return to making food products safely.
Leverage Strain Typing to Improve Your Food Safety Program
Microbial strain typing represents a powerful tool for food manufacturers, helping you get to the bottom of repeat positives once and for all.
By understanding and harnessing this technology, you can improve your ability to detect and mitigate microbial risks, ultimately protecting your consumers and brand. In an industry where consumer trust is paramount, using advanced techniques like microbial strain typing can make all the difference.



